Hyperparameter Optimization Example¶
This Jupyter Notebook is made for illustrating - through a mixture of slides and code in an interactive fashion - the different methods for optimising Hyperparameters for Machine Learning models. First it shows the most naive, manual approach, then grid search, and finally bayesian optimization.
Authors and Date:¶
- Christian Michelsen & Troels Petersen (Niels Bohr Institute)
- 2026-04-22 (latest update)
- Naive, manual approach
- Grid search
- Random search
- Bayesian optimization
- "Full" scan over parameter space
- New methods
- New software
- Focus on the understanding of HPO, not the actual code nor the data!
Nomenclature (i.e. naming scheme)¶
- Machine Learning Model: $\mathcal{A}$
- $N$ hyperparameters
- Domain: $\Lambda_n$
- Hyperparameter configuration space: $\mathbf{\Lambda}=\Lambda_1 \times \Lambda_2 \times \dots \times \Lambda_N $
- Vector of hyperparameters: $\mathbf{\lambda} \in \mathbf{\Lambda}$
- Specific ML model: $\mathcal{A}_\mathbf{\lambda}$
Domain of hyperparameters:¶
- real
- integer
- binary
- categorical
Goal:¶
Given a dataset $\mathcal{D}$, find the vector of HyperParameters $\mathbf{\lambda}^{*}$, which performes "best", i.e. minimises the expected loss function $\mathcal{L}$ for the model $\mathcal{A}_\mathbf{\lambda}$ on the test set of the data $D_\mathrm{test}$:
$$ \mathbf{\lambda}^{*} = \mathop{\mathrm{argmin}}_{\mathbf{\lambda} \in \mathbf{\Lambda}} \mathbb{E}_{D_\mathrm{test} \thicksim \mathcal{D}} \, \left[ \mathbf{V}\left(\mathcal{L}, \mathcal{A}_\mathbf{\lambda}, D_\mathrm{test}\right) \right] $$
In practice we have to approximate the expectation above.
First, we import the modules we want to use:
Important¶
Make sure that you have some additional packages installed:
pip install graphviz
conda install -c conda-forge bayesian-optimization
conda install -c conda-forge optuna
pip install optuna-integration
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn import tree
from sklearn.datasets import load_iris, load_wine
from sklearn.metrics import accuracy_score
from IPython.display import SVG
from graphviz import Source
from IPython.display import display
from ipywidgets import interactive
from sklearn.model_selection import train_test_split, GridSearchCV, RandomizedSearchCV
from scipy.stats import randint, poisson
import warnings
# warnings.filterwarnings('ignore')
We read in the data:
df = pd.read_csv('./data/Pulsar_data.csv')
df.head(10)
| Mean_SNR | STD_SNR | Kurtosis_SNR | Skewness_SNR | Class | |
|---|---|---|---|---|---|
| 0 | 27.555184 | 61.719016 | 2.208808 | 3.662680 | 1 |
| 1 | 1.358696 | 13.079034 | 13.312141 | 212.597029 | 1 |
| 2 | 73.112876 | 62.070220 | 1.268206 | 1.082920 | 1 |
| 3 | 146.568562 | 82.394624 | -0.274902 | -1.121848 | 1 |
| 4 | 6.071070 | 29.760400 | 5.318767 | 28.698048 | 1 |
| 5 | 32.919732 | 65.094197 | 1.605538 | 0.871364 | 1 |
| 6 | 34.101171 | 62.577395 | 1.890020 | 2.572133 | 1 |
| 7 | 50.107860 | 66.321825 | 1.456423 | 1.335182 | 1 |
| 8 | 176.119565 | 59.737720 | -1.785377 | 2.940913 | 1 |
| 9 | 183.622910 | 79.932815 | -1.326647 | 0.346712 | 1 |
We then divide the dataset in input features (X) and target (y):
X = df.drop(columns='Class')
y = df['Class']
feature_names = df.columns.tolist()[:-1]
print(X.shape)
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.20,
random_state=42)
X_train.head(10)
(3278, 4)
| Mean_SNR | STD_SNR | Kurtosis_SNR | Skewness_SNR | |
|---|---|---|---|---|
| 233 | 159.849498 | 76.740010 | -0.575016 | -0.941293 |
| 831 | 4.243311 | 26.746490 | 7.110978 | 52.701218 |
| 2658 | 1.015050 | 10.449662 | 15.593479 | 316.011541 |
| 2495 | 2.235786 | 19.071848 | 9.659137 | 99.294390 |
| 2603 | 2.266722 | 15.512103 | 9.062942 | 99.652157 |
| 111 | 121.404682 | 47.965569 | 0.663053 | 1.203139 |
| 1370 | 35.209866 | 60.573157 | 1.635995 | 1.609377 |
| 1124 | 199.577759 | 58.656643 | -1.862320 | 2.391870 |
| 2170 | 0.663043 | 8.571517 | 23.415092 | 655.614875 |
| 2177 | 3.112876 | 16.855717 | 8.301954 | 90.378150 |
And check out the y values (which turns out to be balanced):
y_train.head(10)
233 1 831 1 2658 0 2495 0 2603 0 111 1 1370 1 1124 1 2170 0 2177 0 Name: Class, dtype: int64
y_train.value_counts()
Class 0 1319 1 1303 Name: count, dtype: int64
Part A: Naïve Approach¶
- Manual configuration
- "Babysitting is also known as Trial & Error or Grad Student Descent in the academic field"
def fit_and_grapth_estimator(estimator):
estimator.fit(X_train, y_train)
accuracy = accuracy_score(y_train, estimator.predict(X_train))
print(f'Training Accuracy: {accuracy:.4f}')
class_names = [str(i) for i in range(y_train.nunique())]
graph = Source(tree.export_graphviz(estimator,
out_file=None,
feature_names=feature_names,
class_names=class_names,
filled = True))
display(SVG(graph.pipe(format='svg')))
return estimator
def plot_tree(max_depth=1, min_samples_leaf=1):
estimator = DecisionTreeClassifier(random_state=42,
max_depth=max_depth,
min_samples_leaf=min_samples_leaf)
return fit_and_grapth_estimator(estimator)
display(interactive(plot_tree,
max_depth=(1, 10, 1),
min_samples_leaf=(1, 100, 1)))
interactive(children=(IntSlider(value=1, description='max_depth', max=10, min=1), IntSlider(value=1, descripti…
(Test this interactively in notebook)
And test this configuration out on the test data:
clf_manual = DecisionTreeClassifier(random_state=42,
max_depth=10,
min_samples_leaf=5)
clf_manual.fit(X_train, y_train)
accuracy_manual = accuracy_score(y_test, clf_manual.predict(X_test))
print(f'Accuracy Manual: {accuracy_manual:.4f}')
Accuracy Manual: 0.8201
Part B: Grid Search¶
Grid Search:
- full factorial design
- Cartesian product
- Curse of dimensionality (grows exponentially)
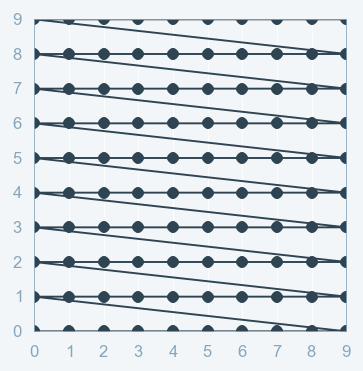
Grid Search with Scikit Learn:
parameters_GridSearch = {'max_depth':[1, 10, 100],
'min_samples_leaf':[1, 10, 100],
}
clf_DecisionTree = DecisionTreeClassifier(random_state=42)
GridSearch = GridSearchCV(clf_DecisionTree,
parameters_GridSearch,
cv=5,
return_train_score=True,
refit=True,
)
GridSearch.fit(X_train, y_train);
GridSearch_results = pd.DataFrame(GridSearch.cv_results_)
print("Grid Search: \tBest parameters: ", GridSearch.best_params_, f", Best scores: {GridSearch.best_score_:.4f}\n")
Grid Search: Best parameters: {'max_depth': 1, 'min_samples_leaf': 1} , Best scores: 0.8551
GridSearch_results
| mean_fit_time | std_fit_time | mean_score_time | std_score_time | param_max_depth | param_min_samples_leaf | params | split0_test_score | split1_test_score | split2_test_score | ... | mean_test_score | std_test_score | rank_test_score | split0_train_score | split1_train_score | split2_train_score | split3_train_score | split4_train_score | mean_train_score | std_train_score | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.001118 | 0.000165 | 0.000350 | 0.000055 | 1 | 1 | {'max_depth': 1, 'min_samples_leaf': 1} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 1 | 0.000961 | 0.000017 | 0.000290 | 0.000020 | 1 | 10 | {'max_depth': 1, 'min_samples_leaf': 10} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 2 | 0.001059 | 0.000052 | 0.000346 | 0.000073 | 1 | 100 | {'max_depth': 1, 'min_samples_leaf': 100} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 3 | 0.004282 | 0.000076 | 0.000301 | 0.000002 | 10 | 1 | {'max_depth': 10, 'min_samples_leaf': 1} | 0.847619 | 0.822857 | 0.858779 | ... | 0.841729 | 0.011991 | 8 | 0.956128 | 0.948021 | 0.954242 | 0.962345 | 0.957102 | 0.955568 | 0.004633 |
| 4 | 0.003857 | 0.000054 | 0.000303 | 0.000010 | 10 | 10 | {'max_depth': 10, 'min_samples_leaf': 10} | 0.845714 | 0.847619 | 0.853053 | ... | 0.846682 | 0.008059 | 6 | 0.896042 | 0.898903 | 0.898475 | 0.893232 | 0.895615 | 0.896453 | 0.002066 |
| 5 | 0.002419 | 0.000064 | 0.000288 | 0.000009 | 10 | 100 | {'max_depth': 10, 'min_samples_leaf': 100} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 6 | 0.004755 | 0.000115 | 0.000305 | 0.000011 | 100 | 1 | {'max_depth': 100, 'min_samples_leaf': 1} | 0.826667 | 0.796190 | 0.841603 | ... | 0.824190 | 0.015056 | 9 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 1.000000 | 0.000000 |
| 7 | 0.003969 | 0.000053 | 0.000305 | 0.000008 | 100 | 10 | {'max_depth': 100, 'min_samples_leaf': 10} | 0.845714 | 0.849524 | 0.854962 | ... | 0.845536 | 0.009138 | 7 | 0.896996 | 0.899857 | 0.898475 | 0.894185 | 0.896568 | 0.897216 | 0.001909 |
| 8 | 0.002447 | 0.000084 | 0.000313 | 0.000031 | 100 | 100 | {'max_depth': 100, 'min_samples_leaf': 100} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
9 rows × 22 columns
clf_GridSearch = GridSearch.best_estimator_
accuracy_GridSearch = accuracy_score(y_test, clf_GridSearch.predict(X_test))
print(f'Accuracy Manual: {accuracy_manual:.4f}')
print(f'Accuracy Grid Search: {accuracy_GridSearch:.4f}')
Accuracy Manual: 0.8201 Accuracy Grid Search: 0.8430
Part C: Random Search¶
- $B$ function evaluations, $N$ hyperparameters, $y$ number of different values:
$$ y_{\mathrm{Grid Search}} = B^{1/N}, \quad y_{\mathrm{Random Search}} = B $$
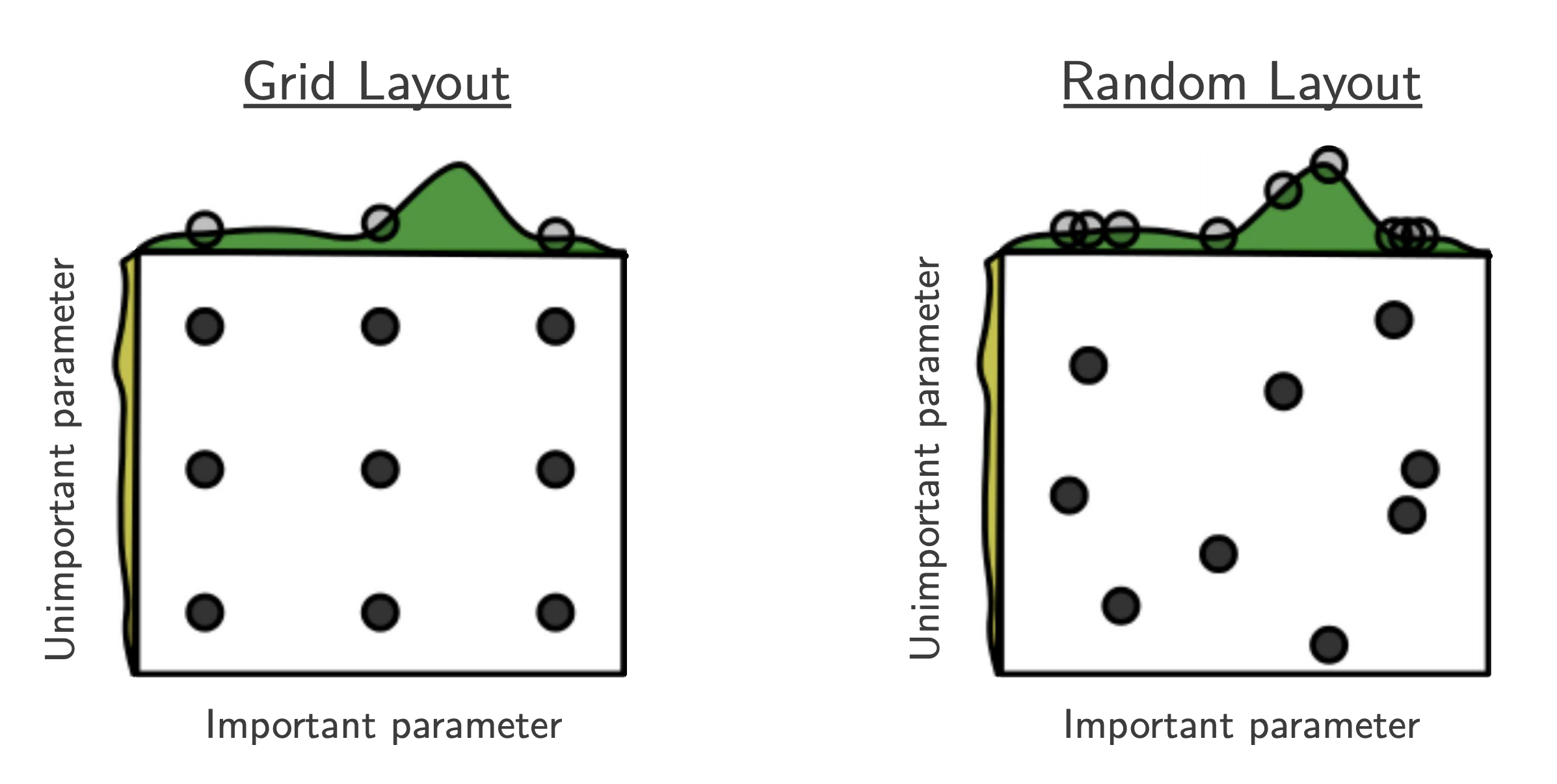
- "This failure of grid search is the rule rather than the exception in high dimensional hyper-parameter optimization" [Bergstra, 2012]
- useful baseline because (almost) no assumptions about the ML algorithm being optimized.
Random Search with Scikit Learn using Scipy Stats as PDFs for the parameters:
# specify parameters and distributions to sample from
parameters_RandomSearch = {'max_depth': poisson(25),
'min_samples_leaf': randint(1, 100)}
# run randomized search
n_iter_search = 9
RandomSearch = RandomizedSearchCV(clf_DecisionTree,
param_distributions=parameters_RandomSearch,
n_iter=n_iter_search,
cv=5,
return_train_score=True,
random_state=42,
)
# fit the random search instance
RandomSearch.fit(X_train, y_train);
RandomSearch_results = pd.DataFrame(RandomSearch.cv_results_)
print("Random Search: \tBest parameters: ", RandomSearch.best_params_, f", Best scores: {RandomSearch.best_score_:.3f}")
Random Search: Best parameters: {'max_depth': 26, 'min_samples_leaf': 83} , Best scores: 0.855
RandomSearch_results.head(10)
| mean_fit_time | std_fit_time | mean_score_time | std_score_time | param_max_depth | param_min_samples_leaf | params | split0_test_score | split1_test_score | split2_test_score | ... | mean_test_score | std_test_score | rank_test_score | split0_train_score | split1_train_score | split2_train_score | split3_train_score | split4_train_score | mean_train_score | std_train_score | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.002836 | 0.000144 | 0.000342 | 0.000047 | 23 | 72 | {'max_depth': 23, 'min_samples_leaf': 72} | 0.849524 | 0.862857 | 0.862595 | ... | 0.854308 | 0.010440 | 7 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 1 | 0.002662 | 0.000060 | 0.000324 | 0.000061 | 26 | 83 | {'max_depth': 26, 'min_samples_leaf': 83} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 2 | 0.003515 | 0.000069 | 0.000300 | 0.000008 | 17 | 24 | {'max_depth': 17, 'min_samples_leaf': 24} | 0.847619 | 0.860952 | 0.868321 | ... | 0.854310 | 0.012162 | 6 | 0.875536 | 0.879351 | 0.878456 | 0.875596 | 0.881792 | 0.878146 | 0.002373 |
| 3 | 0.002520 | 0.000038 | 0.000290 | 0.000006 | 27 | 88 | {'max_depth': 27, 'min_samples_leaf': 88} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 4 | 0.002737 | 0.000029 | 0.000292 | 0.000005 | 31 | 64 | {'max_depth': 31, 'min_samples_leaf': 64} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.861296 | 0.860031 | 0.001460 |
| 5 | 0.002541 | 0.000067 | 0.000301 | 0.000012 | 27 | 89 | {'max_depth': 27, 'min_samples_leaf': 89} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.860343 | 0.859840 | 0.001340 |
| 6 | 0.003101 | 0.000033 | 0.000300 | 0.000016 | 22 | 42 | {'max_depth': 22, 'min_samples_leaf': 42} | 0.849524 | 0.836190 | 0.856870 | ... | 0.845540 | 0.015574 | 9 | 0.866476 | 0.865999 | 0.862726 | 0.865110 | 0.864156 | 0.864893 | 0.001343 |
| 7 | 0.002784 | 0.000057 | 0.000316 | 0.000050 | 21 | 62 | {'max_depth': 21, 'min_samples_leaf': 62} | 0.849524 | 0.840000 | 0.862595 | ... | 0.850500 | 0.009778 | 8 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.861296 | 0.860031 | 0.001460 |
| 8 | 0.002773 | 0.000018 | 0.000307 | 0.000008 | 20 | 64 | {'max_depth': 20, 'min_samples_leaf': 64} | 0.849524 | 0.862857 | 0.862595 | ... | 0.855072 | 0.009121 | 1 | 0.862184 | 0.858369 | 0.858913 | 0.859390 | 0.861296 | 0.860031 | 0.001460 |
9 rows × 22 columns
clf_RandomSearch = RandomSearch.best_estimator_
accuracy_RandomSearch = accuracy_score(y_test, clf_RandomSearch.predict(X_test))
print(f'Accuracy Manual: {accuracy_manual:.4f}')
print(f'Accuracy Grid search: {accuracy_GridSearch:.4f}')
print(f'Accuracy Random Search: {accuracy_RandomSearch:.4f}')
Accuracy Manual: 0.8201 Accuracy Grid search: 0.8430 Accuracy Random Search: 0.8430
Part D: Bayesian Optimization¶
- Expensive black box functions $\Rightarrow$ need of smart guesses
- Probabilistic Surrogate Model (to be fitted)
- Often Gaussian Processes
- Acquisition function
- Exploitation / Exploration
- Cheap to Computer
[Brochu, Cora, de Freitas, 2010]
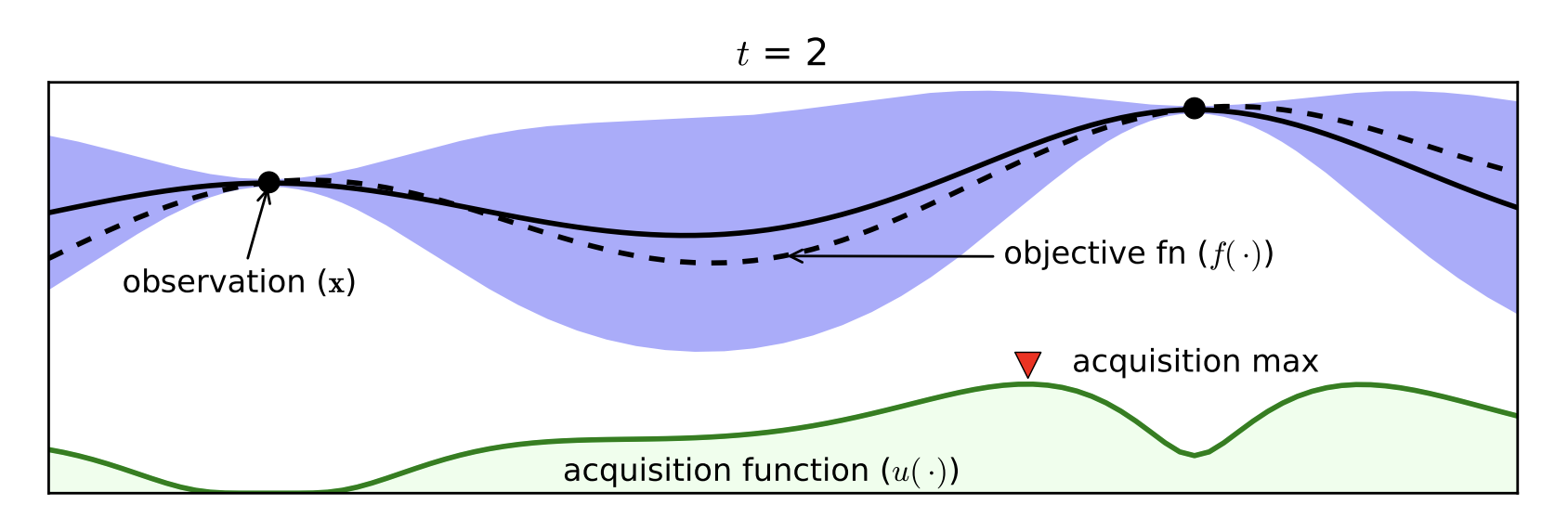
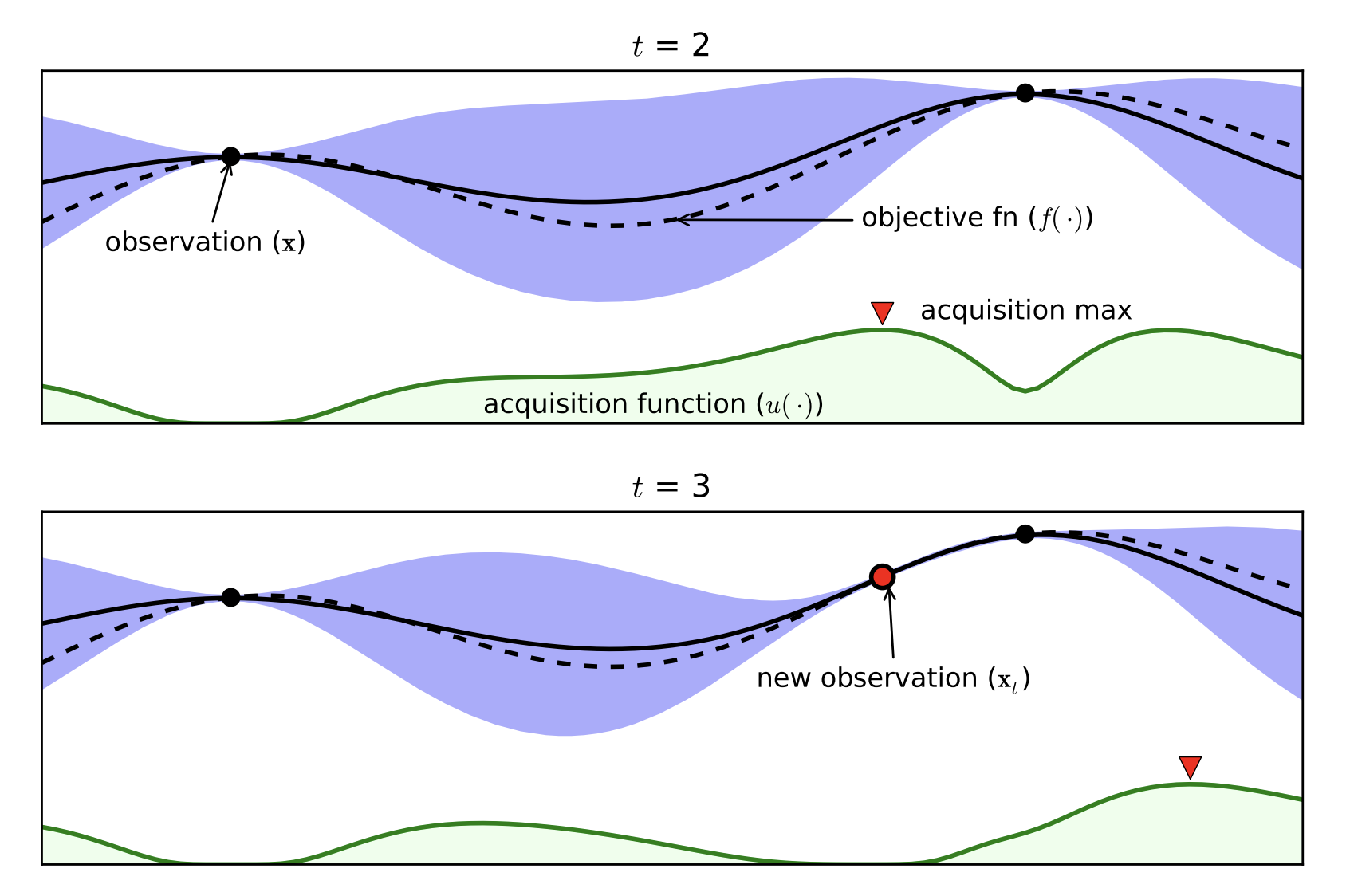
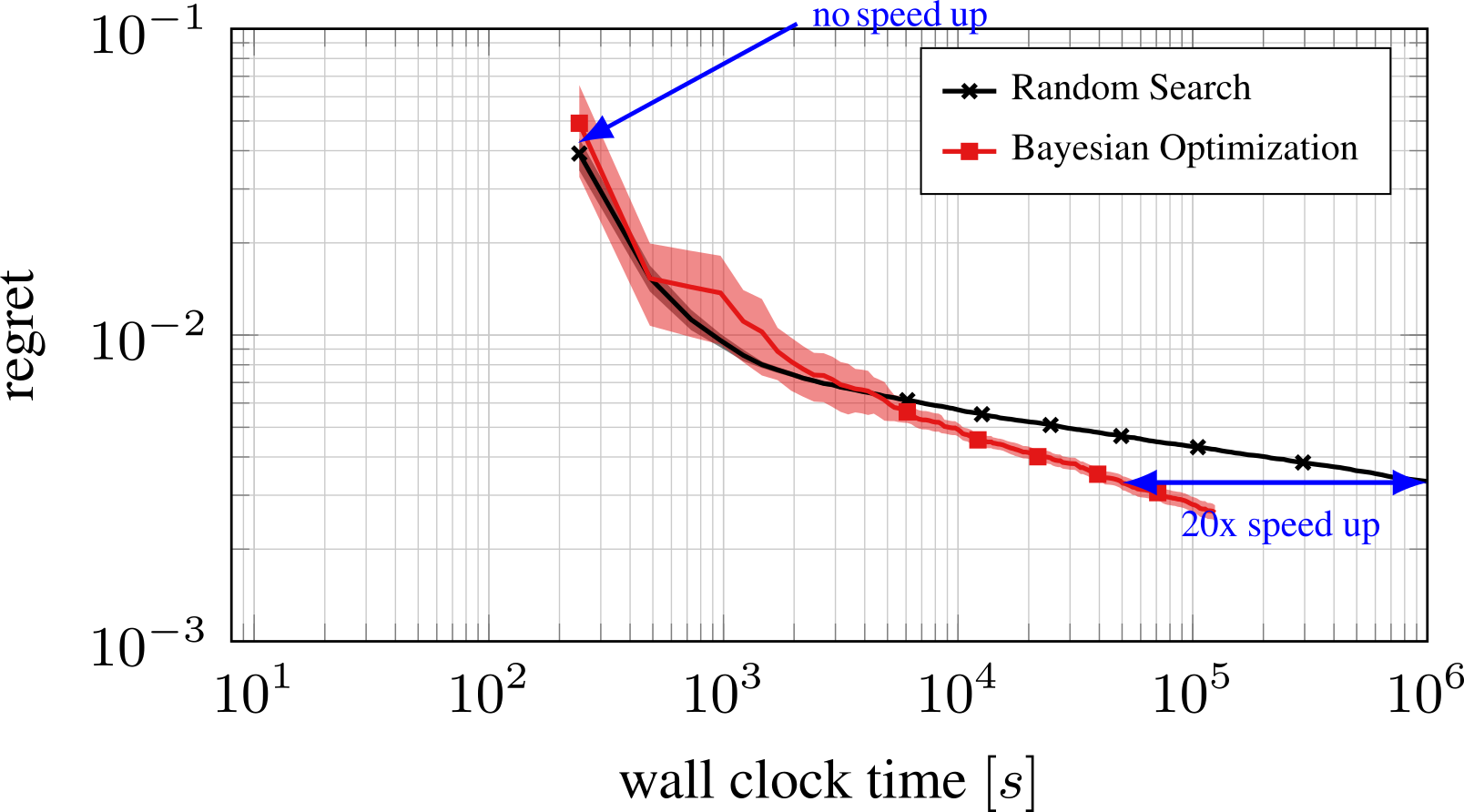
Bayesian Optimization with the Python package BayesianOptimization:
from bayes_opt import BayesianOptimization
from sklearn.model_selection import cross_val_score
def DecisionTree_CrossValidation(max_depth, min_samples_leaf, data, targets):
"""Decision Tree cross validation.
Fits a Decision Tree with the given paramaters to the target
given data, calculated a CV accuracy score and returns the mean.
The goal is to find combinations of max_depth, min_samples_leaf
that maximize the accuracy
"""
estimator = DecisionTreeClassifier(random_state=42,
max_depth=max_depth,
min_samples_leaf=min_samples_leaf)
cval = cross_val_score(estimator, data, targets, scoring='accuracy', cv=5)
return cval.mean()
def optimize_DecisionTree(data, targets, pars, n_iter=5):
"""Apply Bayesian Optimization to Decision Tree parameters."""
def crossval_wrapper(max_depth, min_samples_leaf):
"""Wrapper of Decision Tree cross validation.
Notice how we ensure max_depth, min_samples_leaf
are casted to integer before we pass them along.
"""
return DecisionTree_CrossValidation(max_depth=int(max_depth),
min_samples_leaf=int(min_samples_leaf),
data=data,
targets=targets)
optimizer = BayesianOptimization(f=crossval_wrapper,
pbounds=pars,
random_state=42,
verbose=2)
optimizer.maximize(init_points=4, n_iter=n_iter)
return optimizer
parameters_BayesianOptimization = {"max_depth": (1, 100),
"min_samples_leaf": (1, 100),
}
BayesianOptimization = optimize_DecisionTree(X_train,
y_train,
parameters_BayesianOptimization,
n_iter=5)
print(BayesianOptimization.max)
| iter | target | max_depth | min_sa... |
-------------------------------------------------
| 1 | 0.8550716 | 38.079471 | 95.120716 |
| 2 | 0.8489741 | 73.467400 | 60.267189 |
| 3 | 0.8524020 | 16.445845 | 16.443457 |
| 4 | 0.8550716 | 6.7502776 | 86.751438 |
| 5 | 0.8550716 | 22.279097 | 67.167709 |
| 6 | 0.8550716 | 22.137675 | 65.643624 |
| 7 | 0.8241897 | 100.0 | 1.0 |
| 8 | 0.8550716 | 100.0 | 100.0 |
| 9 | 0.8550716 | 1.0 | 40.766318 |
=================================================
{'target': np.float64(0.8550716103235187), 'params': {'max_depth': np.float64(38.07947176588889), 'min_samples_leaf': np.float64(95.1207163345817)}}
params = BayesianOptimization.max['params']
clf_BO = DecisionTreeClassifier(random_state=42, **params)
clf_BO = clf_BO.fit(X_train, y_train)
accuracy_BayesianOptimization = accuracy_score(y_test, clf_BO.predict(X_test))
print(f'Accuracy Manual: {accuracy_manual:.4f}')
print(f'Accuracy Grid Search: {accuracy_GridSearch:.4f}')
print(f'Accuracy Random Search: {accuracy_RandomSearch:.4f}')
print(f'Accuracy Bayesian Optimization: {accuracy_BayesianOptimization:.4f}')
Accuracy Manual: 0.8201 Accuracy Grid Search: 0.8430 Accuracy Random Search: 0.8430 Accuracy Bayesian Optimization: 0.8430
Alternative dataset: There isn't one perfect hyperparameters optimization algorithm. Try to run this script with another dataset to see how the accuracy results change. For minimal changes with the bjet dataset, replace the line:
df = pd.read_csv('./data/Pulsar_data.csv')
at the beginning of the code with:
import sys
# Linking to the dataset in a parent directory
sys.path.append("../../")
df = pd.DataFrame(np.genfromtxt('../../Week1/AlephBtag_MC_train_Nev5000.csv', names=True))
# The target variable is isb in the other dataset
df['Class'] = data['isb']
# Removing columns we don't want to train on
df = df.drop(columns=['isb', 'nnbjet', 'energy', 'cTheta', 'phi'])
You can also try with the 50000 events dataset by adding a "0" to the filename.
Part D: Full Scan over Parameter Space¶
Only possible in low-dimensional space, slow
max_depth_array = np.arange(1, 30)
min_samples_leaf_array = np.arange(2, 31)
Z = np.zeros((len(max_depth_array), len(min_samples_leaf_array)))
for i, max_depth in enumerate(max_depth_array):
for j, min_samples_leaf in enumerate(min_samples_leaf_array):
clf = DecisionTreeClassifier(random_state=42,
max_depth=max_depth,
min_samples_leaf=
min_samples_leaf)
clf.fit(X_train, y_train)
acc = accuracy_score(y_test, clf.predict(X_test))
Z[i, j] = acc
# Notice: have to transpose Z to match up with imshow
Z = Z.T
Plot the results:
fig, ax = plt.subplots(figsize=(12, 6))
# notice that we are setting the extent and origin keywords
CS = ax.imshow(Z, extent=[1, 30, 2, 31], cmap='viridis', origin='lower')
ax.set(xlabel='max_depth', ylabel='min_samples_leaf')
fig.colorbar(CS);
Sum up:¶
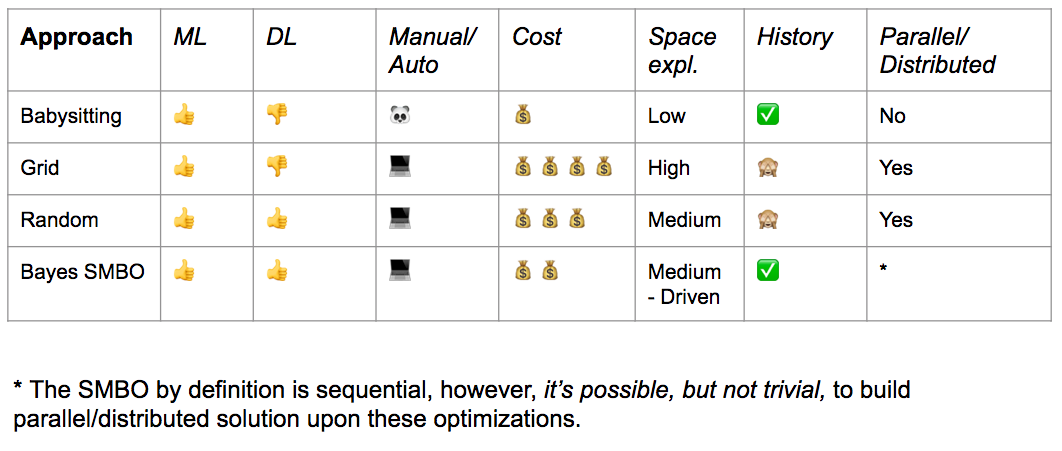
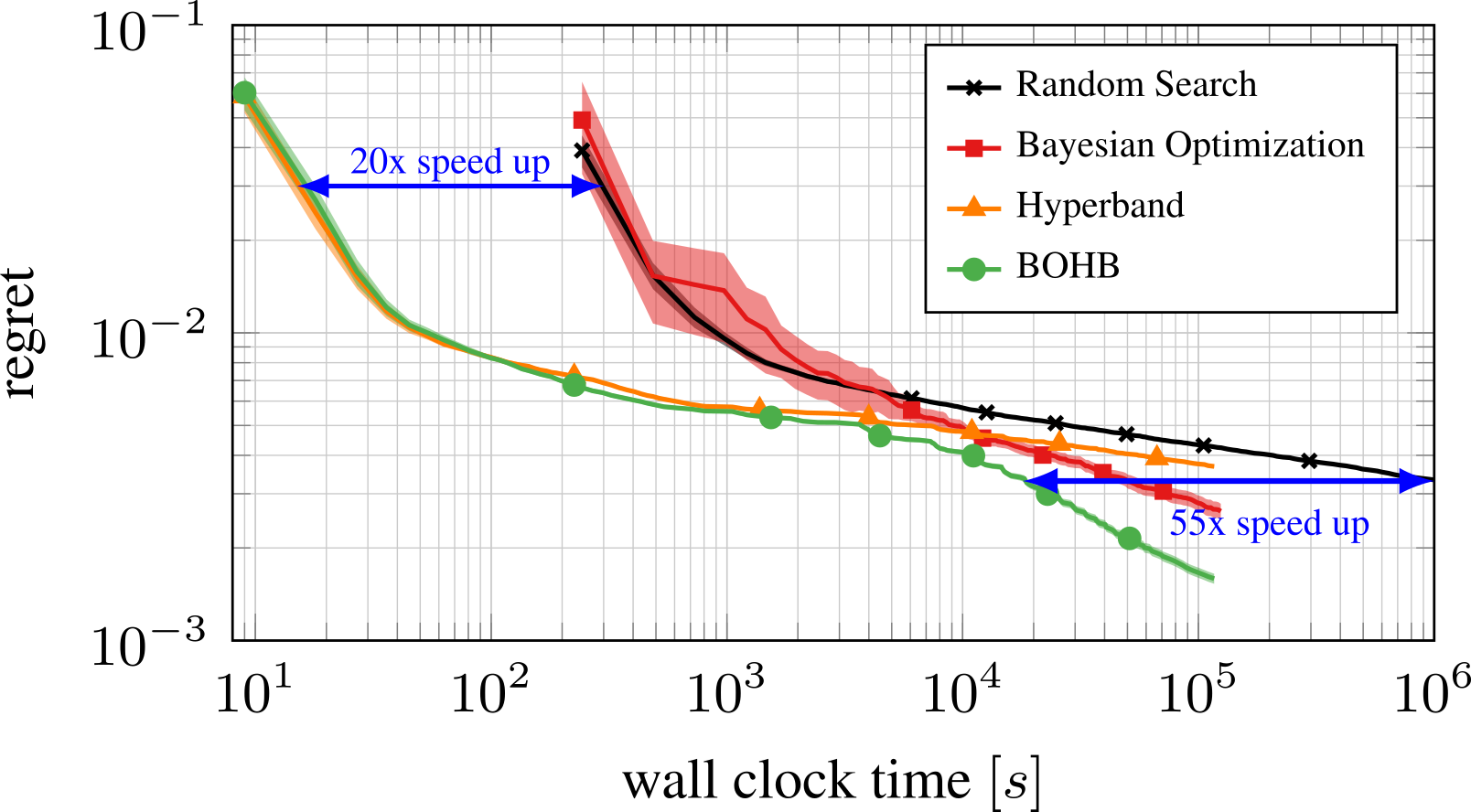
Part E: New Methods¶
Bayesian Optimization meets HyperBand (BOHB)
HyperBand:


import optuna
from optuna.samplers import TPESampler
from optuna.integration import LightGBMPruningCallback
from optuna.pruners import MedianPruner
import lightgbm as lgb
lgb_data_train = lgb.Dataset(X_train, label=y_train);
def objective(trial):
boosting_types = ["gbdt", "rf", "dart"]
boosting_type = trial.suggest_categorical("boosting_type", boosting_types)
params = {
"objective": "binary",
"metric": 'auc',
"boosting": boosting_type,
"max_depth": 5,
"max_depth": trial.suggest_int("max_depth", 2, 63),
"min_child_weight": trial.suggest_float("min_child_weight", 1e-5, 10, log=True),
"scale_pos_weight": trial.suggest_float("scale_pos_weight", 10.0, 30.0),
"bagging_freq": 1, "bagging_fraction": 0.6,
"verbosity": -1
}
N_iterations_max = 10_000
early_stopping_rounds = 10
if boosting_type == "dart":
N_iterations_max = 100
early_stopping_rounds = 0
cv_res = lgb.cv(
params,
lgb_data_train,
num_boost_round=N_iterations_max,
seed=42,
callbacks=[LightGBMPruningCallback(trial, "auc"),lgb.early_stopping(stopping_rounds=early_stopping_rounds),lgb.log_evaluation(period=0)],
)
num_boost_round = len(cv_res["valid auc-mean"])
trial.set_user_attr("num_boost_round", num_boost_round)
return cv_res["valid auc-mean"][-1]
study = optuna.create_study(
direction="maximize",
sampler=TPESampler(seed=42),
pruner=MedianPruner(n_warmup_steps=50),
)
study.optimize(objective, n_trials=100, show_progress_bar=True);
[I 2026-04-22 21:58:04,201] A new study created in memory with name: no-name-fdd2556e-ddc3-440a-87c8-48f07aafc319
0%| | 0/100 [00:00<?, ?it/s]
Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [27] valid's auc: 0.923591 + 0.00625717 [I 2026-04-22 21:58:05,180] Trial 0 finished with value: 0.923591091502377 and parameters: {'boosting_type': 'rf', 'max_depth': 39, 'min_child_weight': 8.632008168602535e-05, 'scale_pos_weight': 13.119890406724053}. Best is trial 0 with value: 0.923591091502377. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [30] valid's auc: 0.922688 + 0.00562831 [I 2026-04-22 21:58:06,231] Trial 1 finished with value: 0.9226879026588939 and parameters: {'boosting_type': 'rf', 'max_depth': 45, 'min_child_weight': 1.3289448722869181e-05, 'scale_pos_weight': 29.398197043239886}. Best is trial 0 with value: 0.923591091502377. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [75] valid's auc: 0.931813 + 0.00638752 [I 2026-04-22 21:58:08,488] Trial 2 finished with value: 0.9318133275583781 and parameters: {'boosting_type': 'gbdt', 'max_depth': 13, 'min_child_weight': 0.0006690421166498799, 'scale_pos_weight': 20.495128632644757}. Best is trial 2 with value: 0.9318133275583781.
/opt/homebrew/anaconda3/envs/appml26/lib/python3.13/site-packages/lightgbm/callback.py:333: UserWarning: Early stopping is not available in dart mode
_log_warning("Early stopping is not available in dart mode")
[I 2026-04-22 21:58:11,283] Trial 3 finished with value: 0.9300085140349988 and parameters: {'boosting_type': 'dart', 'max_depth': 10, 'min_child_weight': 0.0005660670699258885, 'scale_pos_weight': 17.327236865873836}. Best is trial 2 with value: 0.9318133275583781. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [23] valid's auc: 0.925103 + 0.00664956 [I 2026-04-22 21:58:12,142] Trial 4 finished with value: 0.925103266293118 and parameters: {'boosting_type': 'rf', 'max_depth': 33, 'min_child_weight': 0.035849855803404725, 'scale_pos_weight': 10.929008254399955}. Best is trial 2 with value: 0.9318133275583781. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:13,504] Trial 5 pruned. Trial was pruned at iteration 50.
/opt/homebrew/anaconda3/envs/appml26/lib/python3.13/site-packages/lightgbm/callback.py:333: UserWarning: Early stopping is not available in dart mode
_log_warning("Early stopping is not available in dart mode")
[I 2026-04-22 21:58:14,891] Trial 6 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [37] valid's auc: 0.922706 + 0.00507721 [I 2026-04-22 21:58:16,156] Trial 7 finished with value: 0.9227063697984439 and parameters: {'boosting_type': 'rf', 'max_depth': 43, 'min_child_weight': 0.0007417652034871827, 'scale_pos_weight': 20.401360423556216}. Best is trial 2 with value: 0.9318133275583781.
/opt/homebrew/anaconda3/envs/appml26/lib/python3.13/site-packages/lightgbm/callback.py:333: UserWarning: Early stopping is not available in dart mode
_log_warning("Early stopping is not available in dart mode")
[I 2026-04-22 21:58:17,543] Trial 8 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [29] valid's auc: 0.923603 + 0.00649557 [I 2026-04-22 21:58:18,562] Trial 9 finished with value: 0.9236031526982261 and parameters: {'boosting_type': 'rf', 'max_depth': 14, 'min_child_weight': 1.867943489455631e-05, 'scale_pos_weight': 16.506606615265287}. Best is trial 2 with value: 0.9318133275583781. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:18,792] Trial 10 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [80] valid's auc: 0.932005 + 0.00572181 [I 2026-04-22 21:58:21,244] Trial 11 finished with value: 0.9320045589835481 and parameters: {'boosting_type': 'gbdt', 'max_depth': 17, 'min_child_weight': 0.0014474404511749194, 'scale_pos_weight': 17.256467103396812}. Best is trial 11 with value: 0.9320045589835481. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:22,603] Trial 12 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [75] valid's auc: 0.932222 + 0.00647982 [I 2026-04-22 21:58:24,905] Trial 13 finished with value: 0.9322223804893104 and parameters: {'boosting_type': 'gbdt', 'max_depth': 21, 'min_child_weight': 0.009013452524358742, 'scale_pos_weight': 16.365725002496163}. Best is trial 13 with value: 0.9322223804893104. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [80] valid's auc: 0.932962 + 0.00616164 [I 2026-04-22 21:58:27,335] Trial 14 finished with value: 0.9329620531129151 and parameters: {'boosting_type': 'gbdt', 'max_depth': 24, 'min_child_weight': 0.5206341066619696, 'scale_pos_weight': 14.999056028284222}. Best is trial 14 with value: 0.9329620531129151. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [76] valid's auc: 0.932264 + 0.00804279 [I 2026-04-22 21:58:29,666] Trial 15 finished with value: 0.9322644588626462 and parameters: {'boosting_type': 'gbdt', 'max_depth': 27, 'min_child_weight': 0.46818808709793036, 'scale_pos_weight': 14.221884014764331}. Best is trial 14 with value: 0.9329620531129151. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:31,502] Trial 16 pruned. Trial was pruned at iteration 67. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:33,282] Trial 17 pruned. Trial was pruned at iteration 65. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:34,646] Trial 18 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:35,172] Trial 19 pruned. Trial was pruned at iteration 50.
/opt/homebrew/anaconda3/envs/appml26/lib/python3.13/site-packages/lightgbm/callback.py:333: UserWarning: Early stopping is not available in dart mode
_log_warning("Early stopping is not available in dart mode")
[I 2026-04-22 21:58:36,558] Trial 20 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:37,921] Trial 21 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:39,291] Trial 22 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:40,737] Trial 23 pruned. Trial was pruned at iteration 53. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:42,362] Trial 24 pruned. Trial was pruned at iteration 59. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:43,725] Trial 25 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:44,920] Trial 26 pruned. Trial was pruned at iteration 53. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:46,278] Trial 27 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:47,931] Trial 28 pruned. Trial was pruned at iteration 61.
/opt/homebrew/anaconda3/envs/appml26/lib/python3.13/site-packages/lightgbm/callback.py:333: UserWarning: Early stopping is not available in dart mode
_log_warning("Early stopping is not available in dart mode")
[I 2026-04-22 21:58:49,316] Trial 29 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [27] valid's auc: 0.923732 + 0.00629018 [I 2026-04-22 21:58:50,279] Trial 30 finished with value: 0.9237324157136173 and parameters: {'boosting_type': 'rf', 'max_depth': 31, 'min_child_weight': 0.01665309875195129, 'scale_pos_weight': 13.432990381665874}. Best is trial 14 with value: 0.9329620531129151. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:51,633] Trial 31 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:53,019] Trial 32 pruned. Trial was pruned at iteration 51. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:54,369] Trial 33 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:55,712] Trial 34 pruned. Trial was pruned at iteration 50. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:57,316] Trial 35 pruned. Trial was pruned at iteration 59. Training until validation scores don't improve for 10 rounds Early stopping, best iteration is: [38] valid's auc: 0.923515 + 0.00569202 [I 2026-04-22 21:58:58,564] Trial 36 finished with value: 0.9235154085925196 and parameters: {'boosting_type': 'rf', 'max_depth': 9, 'min_child_weight': 0.0004218437241982917, 'scale_pos_weight': 19.39629668550541}. Best is trial 14 with value: 0.9329620531129151. Training until validation scores don't improve for 10 rounds [I 2026-04-22 21:58:59,989] Trial 37 pruned. Trial was pruned at iteration 52.
/opt/homebrew/anaconda3/envs/appml26/lib/python3.13/site-packages/lightgbm/callback.py:333: UserWarning: Early stopping is not available in dart mode
_log_warning("Early stopping is not available in dart mode")
# To see all info at the best trial use:
study.best_trial
# To print metric values for all trials:
study.best_trial.intermediate_values
# To see distributions from which optuna samples parameters:
study.best_trial.distributions
# To simply get the optimized parameters:
study.best_trial.params
Happy HyperParameter Optimisation!¶
...and remember, that this is useful but not essential in this course.